The benzodiazepine data set was generated by doing a PubChem search for the SMARTS query "C2(CN=C(C1=CC(=CC=C1N2[*])[*])C3=CC=CC=C3[*])=[*]" where the [*] are for the R-groups. The query pattern was generated by the PubChem sketcher, and the aromatic form without R-groups is "c1ccc2c(c1)C(=NCCN2)c3ccccc3".
This task will extract a molecule from that data set and depict it with part of the substructure highlighted.
Implementation[]
Read record 3016 from the benzodiazepine SD file. Find all atoms which match the SMARTS "c1ccc2c(c1)C(=NCCN2)c3ccccc3" and highlight them in red. All other atoms must be drawn in black.
The resulting image should be 200x250 pixels and on a white background. The resulting image file should be in PNG (preferred) or GIF format.
OpenEye/Python[]
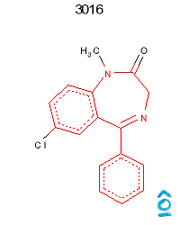
Output from OpenEye/Python
from openeye.oechem import *
from openeye.oedepict import *
# Find the structure matching the given title
def get_molecule_by_title(ifs, title):
for mol in ifs.GetOEGraphMols():
if mol.GetTitle() == title:
return mol
raise KeyError(title)
def do_layout_and_make_view(mol):
# These operations may modify the structure
OEAssignAromaticFlags(mol)
OESetDimensionFromCoords(mol)
OESuppressHydrogens(mol)
OEAddDepictionHydrogens(mol)
OEDepictCoordinates(mol)
OEMDLPerceiveBondStereo(mol)
view = OEDepictView(200, 250)
view.SetSuperAtoms(False)
view.SetAromaticCircles(False)
view.SetAromaticDashes(True)
view.SetColorOnBlack(False)
view.SetBackColor(255, 255, 255)
view.SetForeColor(0, 0, 0)
view.SetShowHydrogens(True)
view.SetDativeBonds(False)
view.SetMolecule(mol)
return view
# Set the color of each atom, as specified
def color_atoms(view, atoms, (r, g, b)):
for atom in atoms:
astyle = view.AStyle(atom.GetIdx())
astyle.r, astyle.g, astyle.b = r, g, b
# Get the right molecule
ifs = oemolistream("benzodiazepine.sdf.gz")
mol = get_molecule_by_title(ifs, "3016")
# prepare it for depiction and make everything black
view = do_layout_and_make_view(mol)
color_atoms(view, mol.GetAtoms(), (0, 0, 0))
# Define the core pattern and make all matches red
core_subsearch = OESubSearch("c1ccc2c(c1)C(=NCCN2)c3ccccc3")
for match in core_subsearch.Match(mol, True):
color_atoms(view, match.GetTargetAtoms(), (255, 0, 0))
# Render to an image and write the result as a GIF
img = OE8BitImage(view.XRange(), view.YRange())
view.RenderImage(img, True, 0, 0)
ofs = oeofstream("highlight_oe.gif")
OEWriteGIF(ofs, img)
OpenBabel/Rubabel[]
mol = Rubabel.foreach("benzodiazepine.sdf.gz").find {|mol| mol.title == "3016" }
mol.highlight_substructure!("c1ccc2c(c1)C(=NCCN2)c3ccccc3").remove_h!
mol.write("3016_highlighted.rubabel.png", u: true)
RDKit/Python[]
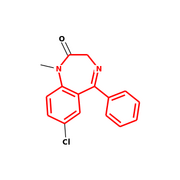
Output from RDKit/Python
from rdkit import Chem
suppl = Chem.SDMolSupplier('benzodiazepine.sdf')
tgt=None
for mol in suppl:
if not mol: continue
if mol.GetProp('_Name')=='3016':
tgt=mol
break
patt = Chem.MolFromSmarts("c1ccc2c(c1)C(=NCCN2)c3ccccc3")
# get the matching atoms:
matching = mol.GetSubstructMatch(patt)
from rdkit.Chem import Draw
# By default the RDKit colors atoms by element in depictions.
# We can turn this off by replacing the element dictionary
# in MolDrawing:
from rdkit.Chem.Draw import MolDrawing
from collections import defaultdict
MolDrawing.elemDict=defaultdict(lambda : (0,0,0))
Draw.MolToFile(tgt,'3016_highlighted.rdkit.png',size=(200,250),highlightAtoms=matching )
Cactvs/Tcl[]
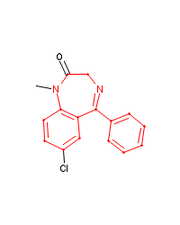
Output from Cactvs
set eh [molfile scan [molfile open benzodiazepine.sdf.gz r hydrogens add] "E_NAME = 3016" ens]
match ss -atomhighlight 1 -bondhighlight 1 c1ccc2c(c1)C(=NCCN2)c3ccccc3 $eh
prop setparam E_GIF height 250 width 200 bgcolor white atomcolor black format png filename 3016.png
ens get $eh E_GIF
The rendering uses the original coordinates from the SD file (unlike the OE example).
Btw: What does "Read record 3016" mean? If this is the record number, and not the record name, the script is:
set eh [molfile read [molfile open benzodiazepine.sdf.gz r record 3016 hydrogens add]]
match ss -atomhighlight 1 -bondhighlight 1 c1ccc2c(c1)C(=NCCN2)c3ccccc3 $eh
prop setparam E_GIF height 250 width 200 bgcolor white atomcolor black format png filename 3016.png
ens get $eh E_GIF
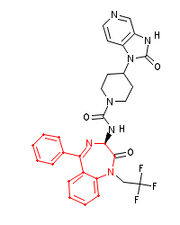
Output from Cactvs
Cactvs/Python[]
First version with ID match:
e=Molfile('benzodiazepine.sdf.gz','r',{'hydrogens':'add'}).scan('E_NAME = 3016','ens')
match('ss','c1ccc2c(c1)C(=NCCN2)c3ccccc3',e,atomhighlight=True,bondhighlight=True)
Prop.Setparam('E_GIF','height',250,'width',200,'bgcolor','white','atomcolor','black','format','png','filename','3016.png')
e.get('E_GIF')
Second version with record selection:
e=Molfile('benzodiazepine.sdf.gz','r',{'record':3016,'hydrogens':'add'}).read()
match('ss','c1ccc2c(c1)C(=NCCN2)c3ccccc3',e,atomhighlight=True,bondhighlight=True)
Prop.Setparam('E_GIF','height',250,'width',200,'bgcolor','white','atomcolor','black','format','png','filename','3016.png')
e.get('E_GIF')
The output is again pixel-identical.
CDK/Java[]
CDK v1.5.12+ provides a new depict API simplifying image generation.
import org.openscience.cdk.CDKConstants;
import org.openscience.cdk.interfaces.*;
import org.openscience.cdk.io.MDLV2000Reader;
import org.openscience.cdk.isomorphism.Pattern;
import org.openscience.cdk.silent.SilentChemObjectBuilder;
import org.openscience.cdk.smiles.smarts.SmartsPattern;
import org.openscience.cdk.tools.manipulator.AtomContainerManipulator;
import java.awt.Color;
import java.io.FileInputStream;
public class Main {
public static void main(String[] args) throws Exception {
String fname = "benzodiazepine.sdf";
String sma = "c1ccc2c(c1)C(=NCCN2)c3ccccc3";
IChemObjectBuilder bldr = SilentChemObjectBuilder.getInstance();
IAtomContainer mol = null;
// XXX: use try-with-resources WIKIA doesn't syntax color if I do
FileInputStream in = new FileInputStream(fname);
MDLV2000Reader mdlr = new MDLV2000Reader(in);
try {
while ((mol = mdlr.read(bldr.newInstance(IAtomContainer.class))) != null) {
if (mol.getProperty(CDKConstants.TITLE).equals("3016")) {
break;
}
}
} finally {
mdlr.close();
}
AtomContainerManipulator.suppressHydrogens(mol);
Pattern ptrn = SmartsPattern.create(sma, bldr);
Iterable<IChemObject> hits = ptrn.matchAll(mol)
.uniqueAtoms()
.toChemObjects();
DepictionGenerator dptgen = new DepictionGenerator();
dptgen.withSize(200, 250)
.withHighlight(hits, Color.RED);
dptgen.depict(mol)
.writeTo("~/3016.png");
}
}
CDK/Groovy[]
Output from CDK/Groovy
import java.util.List;
import java.awt.*;
import java.awt.image.*;
import java.util.zip.GZIPInputStream;
import javax.imageio.*;
import org.openscience.cdk.*;
import org.openscience.cdk.interfaces.*;
import org.openscience.cdk.io.*;
import org.openscience.cdk.io.iterator.*;
import org.openscience.cdk.layout.*;
import org.openscience.cdk.renderer.*;
import org.openscience.cdk.renderer.font.*;
import org.openscience.cdk.renderer.generators.*;
import org.openscience.cdk.renderer.visitor.*;
import org.openscience.cdk.renderer.generators.BasicSceneGenerator.Margin;
import org.openscience.cdk.renderer.generators.BasicSceneGenerator.ZoomFactor;
import org.openscience.cdk.silent.*;
import org.openscience.cdk.smiles.smarts.*;
import org.openscience.cdk.templates.*;
import org.openscience.cdk.tools.manipulator.*;
int WIDTH = 250;
int HEIGHT = 200;
// the draw area and the image should be the same size
Rectangle drawArea = new Rectangle(WIDTH, HEIGHT);
Image image = new BufferedImage(
WIDTH, HEIGHT, BufferedImage.TYPE_INT_RGB
);
iterator = new IteratingMDLReader(
new GZIPInputStream(
new File("ctr/benzodiazepine.sdf.gz")
.newInputStream()
),
SilentChemObjectBuilder.getInstance()
)
iterator.setReaderMode(
IChemObjectReader.Mode.STRICT
)
compound3016 = null
while (iterator.hasNext() && compound3016 == null) {
mol = iterator.next()
if ("3016".equals(mol.getProperty(CDKConstants.TITLE)))
compound3016 = mol
}
compound3016 = AtomContainerManipulator.removeHydrogens(compound3016)
StructureDiagramGenerator sdg = new StructureDiagramGenerator();
sdg.setMolecule(compound3016);
sdg.generateCoordinates();
compound3016 = sdg.getMolecule();
// generators make the image elements
List<IGenerator> generators =
new ArrayList<IGenerator>();
generators.add(new BasicSceneGenerator());
generators.add(new ExternalHighlightGenerator());
generators.add(new BasicBondGenerator());
generators.add(new BasicAtomGenerator());
selection = new AtomContainer();
querytool = new SMARTSQueryTool("c1ccc2c(c1)C(=NCCN2)c3ccccc3");
querytool.matches(compound3016);
if (querytool.countMatches() > 0) {
mappings = querytool.getUniqueMatchingAtoms()
mapping = mappings.get(0)
for (int i=0; i<mapping.size(); i++) {
selection.addAtom(
compound3016.getAtom(mapping.get(i))
)
}
}
// the renderer needs to have a toolkit-specific font manager
AtomContainerRenderer renderer =
new AtomContainerRenderer(generators, new AWTFontManager());
// the call to 'setup' only needs to be done on the first paint
renderer.setup(compound3016, drawArea);
model = renderer.getRenderer2DModel();
model.set(ZoomFactor.class, (double)0.5);
model.set(
ExternalHighlightGenerator
.ExternalHighlightDistance.class,
(double)18
);
model.set(
RendererModel.ExternalHighlightColor.class,
Color.red
);
model.setExternalSelectedPart(selection);
// paint the background
Graphics2D g2 = (Graphics2D)image.getGraphics();
g2.setColor(Color.WHITE);
g2.fillRect(0, 0, WIDTH, HEIGHT);
// the paint method also needs a toolkit-specific renderer
renderer.paint(compound3016, new AWTDrawVisitor(g2));
ImageIO.write(
(RenderedImage)image, "PNG",
new File("CTR7.png")
);
Indigo/Python[]
Output from Indigo/Python
from indigo import Indigo
from indigo_renderer import IndigoRenderer
indigo = Indigo()
renderer = IndigoRenderer(indigo)
tgt=None
for mol in indigo.iterateSDFile("benzodiazepine.sdf"):
if not mol:
continue
if mol.name()=='3016':
tgt=mol
break
query = indigo.loadQueryMolecule("c1ccc2c(c1)C(=NCCN2)c3ccccc3")
match = indigo.substructureMatcher(tgt).match(query)
if match:
matching = match.highlightedTarget()
matching.foldHydrogens() # Indigo does not change the representation of the hydrogens automatically
matching.layout() # if not called, will be done automatically by the renderer
indigo.setOption("render-output-format", "png")
indigo.setOption("render-image-size", 200, 250)
indigo.setOption("render-background-color", 1.0, 1.0, 1.0)
renderer.renderToFile(matching, "3016_highlighted.indigo.png")